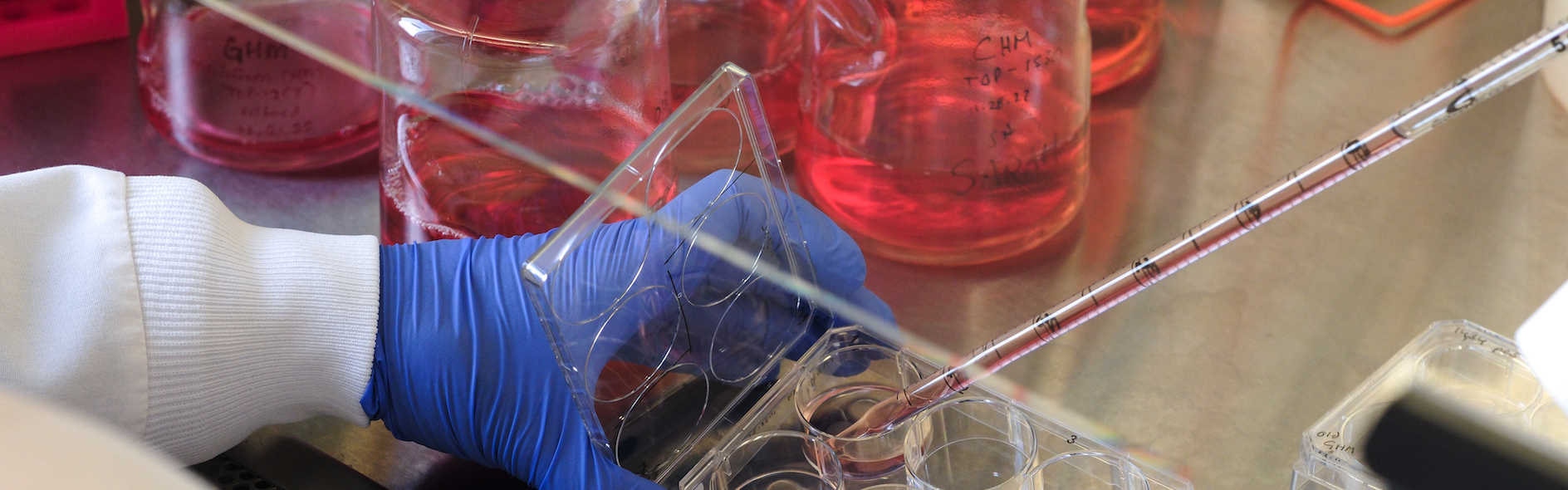
The EIPM develops organoids, also known as patient-derived ex-vivo models, used to study disease progression and develop effective drug treatments. These miniature three-dimensional cellular structures are grown from a patient’s tumor sample and are highly useful in studying how different cancers develop, change, and might respond to various treatment options.
Advancing Treatment for Patients
For patients with advanced disease, organoids can serve as an ideal platform to enable discoveries of novel therapeutic approaches that can be assessed in clinical trials and provide personalized therapeutic options for individual patients where standard clinical options have been exhausted.
The EIPM has been at the forefront of developing organoid technology that will advance science and speed new treatments to patients.
Our platform is unique because we have advanced the technology to derive organoids from many different types of cancer, unlike many other institutions around the globe.
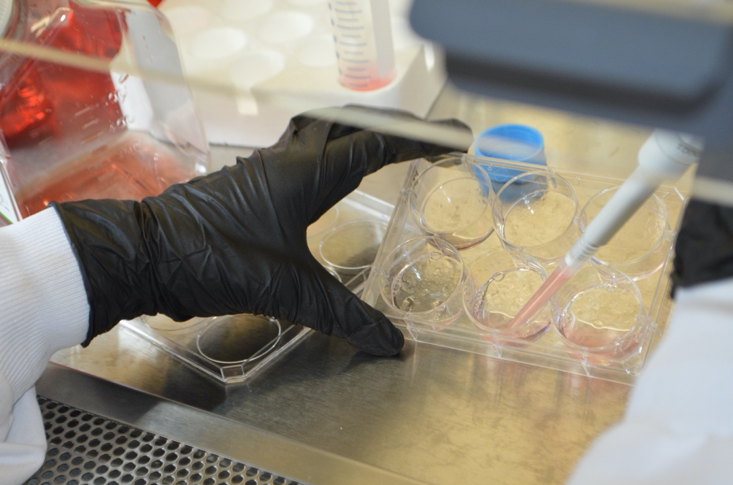
Organoid cell culture in progress.
Workflow
The EIPM team develops organoids from metastatic and primary anatomic sites, obtained through biopsies, surgical resections, and rapid autopsy procedures.
Our clinician colleagues send a tiny biological sample from a cancer patient to the EIPM research lab, where technicians grow dozens or even hundreds of exact organoid replicas of the tumor.
A critical step in successfully generating patient-derived organoids is validating their biological similarity with the primary tumor. This is accomplished through the Englander Institute's Next Generation Sequencing platform.
Following sequencing, investigators are able to characterize each patient-derived organoid with genomic results, RNA-seq, and histopathology. Validation of these models is achieved by comparison of DNA mutational profiles (ploidy, concordance, correlation of variant allele frequencies, mutational signatures) as well as RNA sequencing data through cluster analysis of the original tumor in comparison with the organoids.
Models Available
We now have >250 pan cancer models in our organoid repository at various stages of characterization. Approximately 150 have been characterized via sequencing and/or pathology review and are now available for research use.


